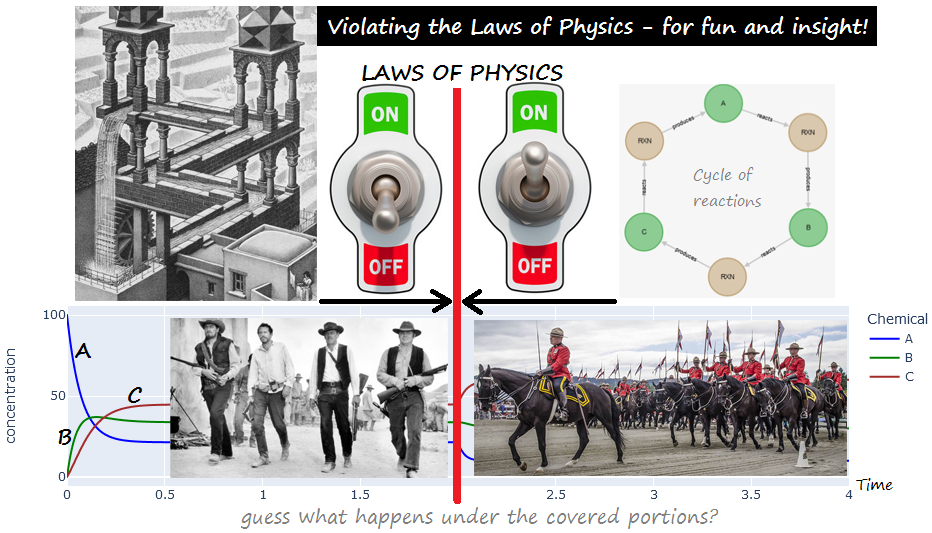
Violating the Laws of Physics for Fun and Insight!¶
A cascade of reactions A <-> B <-> C , mostly in the forward direction¶
PART 1 : the above, together with a PHYSICALLY-IMPOSSIBLE "closing" of the cycle with :¶
C <-> A, ALSO mostly in the forward direction (never mind the laws of thermodymics)!¶
PART 2 : restoring the law of physics (by letting C <-> A adjust its kinetics based on the energy difference.)¶
All 1st-order kinetics.
LAST REVISED: May 25, 2023
In [1]:
import set_path # Importing this module will add the project's home directory to sys.path
Added 'D:\Docs\- MY CODE\BioSimulations\life123-Win7' to sys.path
In [2]:
from experiments.get_notebook_info import get_notebook_basename
from src.modules.reactions.reaction_data import ReactionData as chem
from src.modules.reactions.reaction_dynamics import ReactionDynamics
from src.modules.numerical.numerical import Numerical as num
import numpy as np
import plotly.express as px
import plotly.graph_objects as go
from src.modules.visualization.graphic_log import GraphicLog
In [3]:
# Initialize the HTML logging
log_file = get_notebook_basename() + ".log.htm" # Use the notebook base filename for the log file
# Set up the use of some specified graphic (Vue) components
GraphicLog.config(filename=log_file,
components=["vue_cytoscape_1"],
extra_js="https://cdnjs.cloudflare.com/ajax/libs/cytoscape/3.21.2/cytoscape.umd.js")
-> Output will be LOGGED into the file 'impossible_1.log.htm'
Initialize the system¶
In [4]:
# Initialize the system
chem_data = chem(names=["A", "B", "C"])
# Reaction A <-> B, mostly in forward direction (favored energetically)
# Note: all reactions in this experiment have 1st-order kinetics for all species
chem_data.add_reaction(reactants="A", products="B",
forward_rate=9., reverse_rate=3.)
# Reaction B <-> C, also favored energetically
chem_data.add_reaction(reactants="B", products="C",
forward_rate=8., reverse_rate=4.)
Part 1 - "Turning off the Laws of Physics"!¶
In [5]:
# LET'S VIOLATE THE LAWS OF PHYSICS!
# Reaction C <-> A, also mostly in forward direction - MAGICALLY GOING "UPSTREAM" from C, to the higher-energy level of "A"
chem_data.add_reaction(reactants="C" , products="A",
forward_rate=3., reverse_rate=2.) # PHYSICALLY IMPOSSIBLE! Future versions of Life123 may flag this!
chem_data.describe_reactions()
# Send the plot of the reaction network to the HTML log file
graph_data = chem_data.prepare_graph_network()
GraphicLog.export_plot(graph_data, "vue_cytoscape_1")
Number of reactions: 3 (at temp. 25 C) 0: A <-> B (kF = 9 / kR = 3 / Delta_G = -2,723.41 / K = 3) | 1st order in all reactants & products 1: B <-> C (kF = 8 / kR = 4 / Delta_G = -1,718.28 / K = 2) | 1st order in all reactants & products 2: C <-> A (kF = 3 / kR = 2 / Delta_G = -1,005.13 / K = 1.5) | 1st order in all reactants & products [GRAPHIC ELEMENT SENT TO LOG FILE `impossible_1.log.htm`]
Notice the absurdity of the energy levels always going down, throughout the cycle (like in an Escher painting!)¶

Set the initial concentrations of all the chemicals¶
In [6]:
initial_conc = {"A": 100., "B": 0., "C": 0.}
initial_conc
Out[6]:
{'A': 100.0, 'B': 0.0, 'C': 0.0}
In [7]:
dynamics = ReactionDynamics(reaction_data=chem_data)
dynamics.set_conc(conc=initial_conc, snapshot=True)
dynamics.describe_state()
SYSTEM STATE at Time t = 0: 3 species: Species 0 (A). Conc: 100.0 Species 1 (B). Conc: 0.0 Species 2 (C). Conc: 0.0
In [8]:
dynamics.set_diagnostics() # To save diagnostic information about the call to single_compartment_react()
# All of these settings are currently close to the default values... but subject to change; set for repeatability
dynamics.set_thresholds(norm="norm_A", low=0.5, high=0.8, abort=1.44)
dynamics.set_thresholds(norm="norm_B", low=0.08, high=0.5, abort=1.5)
dynamics.set_step_factors(upshift=1.1, downshift=0.4, abort=0.3)
dynamics.set_error_step_factor(0.2)
dynamics.single_compartment_react(initial_step=0.001, target_end_time=2.0,
variable_steps=True, explain_variable_steps=False)
90 total step(s) taken
In [9]:
dynamics.plot_curves()
In [10]:
# dynamics.explain_time_advance()
# dynamics.get_history()
It might look like an equilibrium has been reached. But NOT! Verify the LACK of final equilibrium state:¶
In [11]:
dynamics.is_in_equilibrium()
A <-> B
Final concentrations: [B] = 33.81 ; [A] = 21.43
1. Ratio of reactant/product concentrations, adjusted for reaction orders: 1.57778
Formula used: [B] / [A]
2. Ratio of forward/reverse reaction rates: 3.0
Discrepancy between the two values: 47.41 %
Reaction is NOT in equilibrium (not within 1% tolerance)
B <-> C
Final concentrations: [C] = 44.76 ; [B] = 33.81
1. Ratio of reactant/product concentrations, adjusted for reaction orders: 1.32394
Formula used: [C] / [B]
2. Ratio of forward/reverse reaction rates: 2.0
Discrepancy between the two values: 33.8 %
Reaction is NOT in equilibrium (not within 1% tolerance)
C <-> A
Final concentrations: [A] = 21.43 ; [C] = 44.76
1. Ratio of reactant/product concentrations, adjusted for reaction orders: 0.478723
Formula used: [A] / [C]
2. Ratio of forward/reverse reaction rates: 1.5
Discrepancy between the two values: 68.09 %
Reaction is NOT in equilibrium (not within 1% tolerance)
Out[11]:
{False: [0, 1, 2]}
Not surprisingly, none of the reactions of this physically-impossible hypothetical system are in equilibrium¶
Even though the concentrations don't change, it's NOT from equilibrium in the reactions - but rather from a balancing out of consuming and replenishing across reactions.¶
Consider, for example, the concentrations of A at the end time, and contributions to its change ("Delta A") from individual reactions affecting A, as available from the diagnostic data:¶
In [12]:
dynamics.get_diagnostic_rxn_data(rxn_index=0, tail=1)
Reaction: A <-> B
Out[12]:
| START_TIME | Delta A | Delta B | Delta C | time_step | caption | |
|---|---|---|---|---|---|---|
| 89 | 1.826083 | -16.303235 | 16.303235 | 0.0 | 0.178317 |
In [13]:
dynamics.get_diagnostic_rxn_data(rxn_index=2, tail=1)
Reaction: C <-> A
Out[13]:
| START_TIME | Delta A | Delta B | Delta C | time_step | caption | |
|---|---|---|---|---|---|---|
| 89 | 1.826083 | 16.303235 | 0.0 | -16.303235 | 0.178317 |
Looking at the last row from each of the 2 dataframes above, one case see that, at every reaction cycle, [A] gets reduced by 0.914286 by the reaction A <-> B, while simultaneously getting increased by the SAME amount by the (fictional) reaction C <-> A.¶
Hence, the concentration of A remains constant - but none of the reactions is in equilibrium!¶
In [ ]:
PART 2 - Let's restore the Laws of Physics!¶
In [14]:
chem_data.describe_reactions()
Number of reactions: 3 (at temp. 25 C) 0: A <-> B (kF = 9 / kR = 3 / Delta_G = -2,723.41 / K = 3) | 1st order in all reactants & products 1: B <-> C (kF = 8 / kR = 4 / Delta_G = -1,718.28 / K = 2) | 1st order in all reactants & products 2: C <-> A (kF = 3 / kR = 2 / Delta_G = -1,005.13 / K = 1.5) | 1st order in all reactants & products
In [15]:
dynamics.clear_reactions() # Let's start over with the reactions (without affecting the data from the reactions)
In [16]:
# For the reactions A <-> B, and B <-> C, everything is being restored to the way it was before
chem_data.add_reaction(reactants="A", products="B",
forward_rate=9., reverse_rate=3.)
# Reaction , also favored energetically
chem_data.add_reaction(reactants="B", products="C",
forward_rate=8., reverse_rate=4.)
In [17]:
chem_data.describe_reactions()
Number of reactions: 2 (at temp. 25 C) 0: A <-> B (kF = 9 / kR = 3 / Delta_G = -2,723.41 / K = 3) | 1st order in all reactants & products 1: B <-> C (kF = 8 / kR = 4 / Delta_G = -1,718.28 / K = 2) | 1st order in all reactants & products
In [18]:
# But for the reaction C <-> A, this time we'll "bend the knee" to the laws of thermodynamics!
# We'll use the same forward rate as before, but we'll let the reverse rate be picked by the system,
# based of thermodynamic data consistent with the previous 2 reactions : i.e. an energy difference of -(-2,723.41 - 1,718.28) = +4,441.69 (reflecting the
# "going uphill energetically" from C to A
chem_data.add_reaction(reactants="C" , products="A",
forward_rate=3., Delta_G=4441.69) # Notice the positive Delta G: we're going from "C", to the higher-energy level of "A"
In [19]:
chem_data.describe_reactions()
Number of reactions: 3 (at temp. 25 C) 0: A <-> B (kF = 9 / kR = 3 / Delta_G = -2,723.41 / K = 3) | 1st order in all reactants & products 1: B <-> C (kF = 8 / kR = 4 / Delta_G = -1,718.28 / K = 2) | 1st order in all reactants & products 2: C <-> A (kF = 3 / kR = 18 / Delta_G = 4,441.69 / K = 0.166667) | 1st order in all reactants & products
Now, let's continue with this "legit" set of reactions, from where we left off in our fantasy world at time t=2:¶
In [20]:
dynamics.single_compartment_react(initial_step=0.01, target_end_time=4.0,
variable_steps=True, explain_variable_steps=False)
#dynamics.explain_time_advance()
#dynamics.get_history()
INFO: the tentative time step (0.01) leads to a least one norm value > its ABORT threshold:
-> will backtrack, and re-do step with a SMALLER delta time, multiplied by 0.3 (set to 0.003) [Step started at t=2.0044, and will rewind there]
45 total step(s) taken
In [21]:
fig0 = dynamics.plot_curves(suppress=True) # Prepare, but don't show, the main plot
In [22]:
# Add a second plot, with a vertical gray line at t=2
fig1 = px.line(x=[2,2], y=[0,100], color_discrete_sequence = ['gray'])
# Combine the plots, and display them
all_fig = go.Figure(data=fig0.data + fig1.data, layout = fig0.layout) # Note that the + is concatenating lists
all_fig.update_layout(title="On the left of vertical gray line: FICTIONAL world; on the right: REAL world!")
all_fig.show()
Notice how [A] drops at time t=2, when we re-enact the Laws of Physics, because A no longer receives the extra boost from the previous mostly-forward (and thus physically-impossible given the unfavorable energy levels!) reaction C <-> A.¶
Back to the real world, that (energetically unfavored) reaction now mostly goes IN REVERSE; hence, the boost in [C] as well¶
Now, we have a REAL equilibrium!¶
In [23]:
dynamics.is_in_equilibrium()
A <-> B
Final concentrations: [B] = 30 ; [A] = 10
1. Ratio of reactant/product concentrations, adjusted for reaction orders: 3
Formula used: [B] / [A]
2. Ratio of forward/reverse reaction rates: 3.0
Discrepancy between the two values: 3.048e-05 %
Reaction IS in equilibrium (within 1% tolerance)
B <-> C
Final concentrations: [C] = 60 ; [B] = 30
1. Ratio of reactant/product concentrations, adjusted for reaction orders: 2
Formula used: [C] / [B]
2. Ratio of forward/reverse reaction rates: 2.0
Discrepancy between the two values: 3.551e-05 %
Reaction IS in equilibrium (within 1% tolerance)
C <-> A
Final concentrations: [A] = 10 ; [C] = 60
1. Ratio of reactant/product concentrations, adjusted for reaction orders: 0.166667
Formula used: [A] / [C]
2. Ratio of forward/reverse reaction rates: 0.16666696258266173
Discrepancy between the two values: 0.0001725 %
Reaction IS in equilibrium (within 1% tolerance)
Out[23]:
True
The fact that individual reactions are now in actual, real equilibrium, can be easily seen from the last rows in the diagnostic data. Notice all the delta-concentration values at the final times are virtually zero:¶
In [24]:
dynamics.get_diagnostic_rxn_data(rxn_index=0, tail=1)
Reaction: A <-> B
Out[24]:
| START_TIME | Delta A | Delta B | Delta C | time_step | caption | |
|---|---|---|---|---|---|---|
| 135 | 3.962322 | -0.000021 | 0.000021 | 0.0 | 0.198792 |
In [25]:
dynamics.get_diagnostic_rxn_data(rxn_index=1, tail=1)
Reaction: B <-> C
Out[25]:
| START_TIME | Delta A | Delta B | Delta C | time_step | caption | |
|---|---|---|---|---|---|---|
| 135 | 3.962322 | 0.0 | -0.000017 | 0.000017 | 0.198792 |
In [26]:
dynamics.get_diagnostic_rxn_data(rxn_index=2, tail=1)
Reaction: C <-> A
Out[26]:
| START_TIME | Delta A | Delta B | Delta C | time_step | caption | |
|---|---|---|---|---|---|---|
| 135 | 3.962322 | 0.000008 | 0.0 | -0.000008 | 0.198792 |
In [ ]: