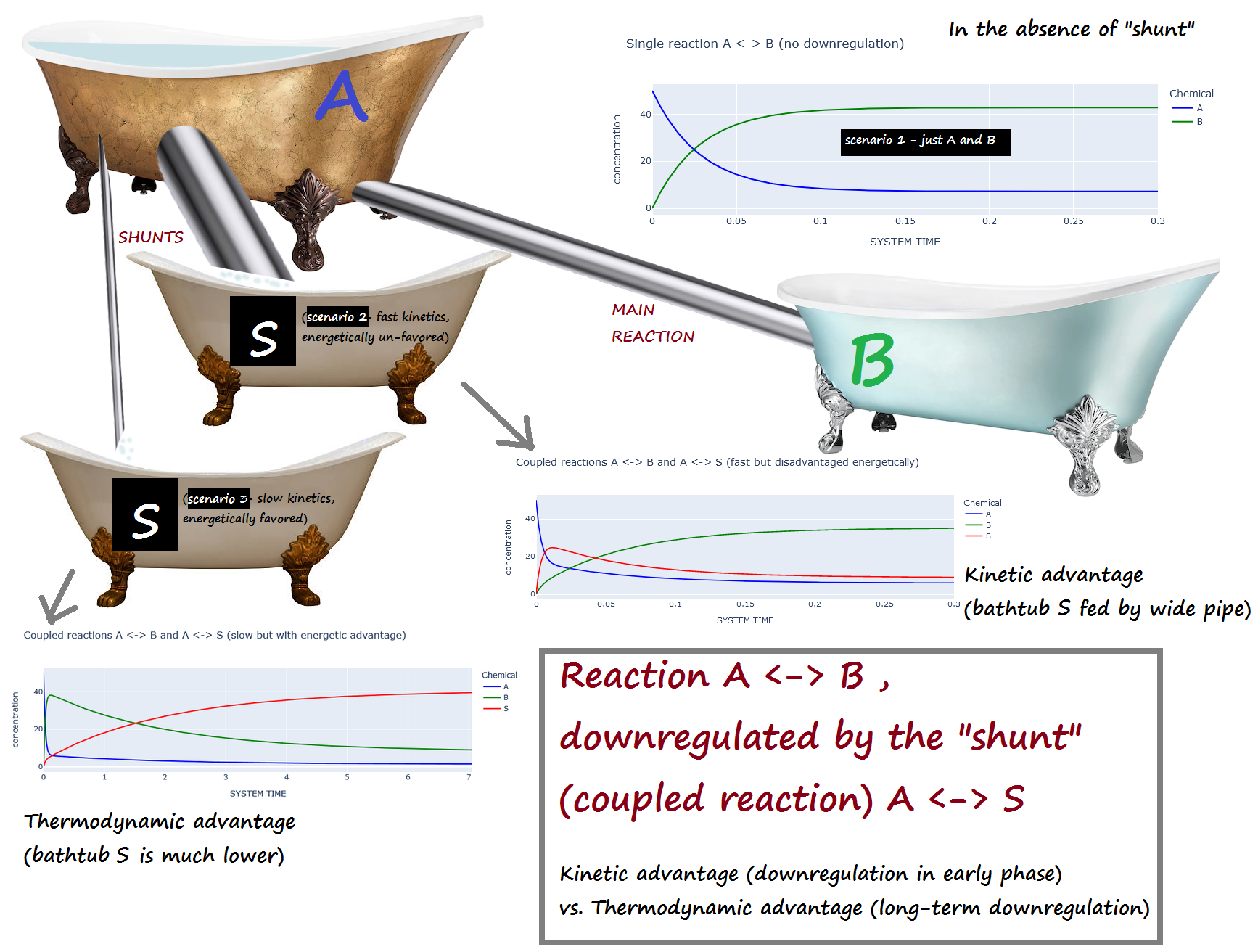
A <-> B , downregulated by the "shunt" (coupled reaction) A <-> S¶
Kinetic advantage (downregulation in early phase) vs. Thermodynamic advantage (long-term downregulation)¶
Scenario 1 : No downregulation on A <-> B
Scenario 2 : The shunt (A <-> S) has a kinetic advantage but thermodynamic DIS-advantage compared to A <-> B
(i.e. A <-> S is fast, but energetically unfavored)
Scenario 3 : The shunt (A <-> S) is has a kinetic DIS-advantage but a thermodynamic advantage compared to A <-> B
(i.e. A <-> S is slow, but energetically favored)
All reactions 1st order, mostly forward. Taken to equilibrium.
LAST REVISED: May 23, 2023
Bathtub analogy:¶
A is initially full, while B and S are empty.
If the "shunt" S is present, scenario 2 corresponds to a large pipe and a small elevation change...
while scenario 3 corresponds to a narrow pipe and a large elevation change.
import set_path # Importing this module will add the project's home directory to sys.path
Added 'D:\Docs\- MY CODE\BioSimulations\life123-Win7' to sys.path
from experiments.get_notebook_info import get_notebook_basename
from src.modules.reactions.reaction_data import ReactionData as chem
from src.modules.reactions.reaction_dynamics import ReactionDynamics
import numpy as np
import plotly.express as px
from src.modules.visualization.graphic_log import GraphicLog
# Initialize the HTML logging (for the graphics)
log_file = get_notebook_basename() + ".log.htm" # Use the notebook base filename for the log file
# Set up the use of some specified graphic (Vue) components
GraphicLog.config(filename=log_file,
components=["vue_cytoscape_1"],
extra_js="https://cdnjs.cloudflare.com/ajax/libs/cytoscape/3.21.2/cytoscape.umd.js")
-> Output will be LOGGED into the file 'down_regulate_1.log.htm'
Scenario 1: A <-> B in the absence of the 2nd reaction¶
Initialize the System¶
Specify the chemicals and the reaction
# Specify the chemicals
chem_data = chem(names=["A", "B"])
# Reaction A <-> B
chem_data.add_reaction(reactants=["A"], products=["B"],
forward_rate=30., reverse_rate=5.)
chem_data.describe_reactions()
Number of reactions: 1 (at temp. 25 C) 0: A <-> B (kF = 30 / kR = 5 / Delta_G = -4,441.69 / K = 6) | 1st order in all reactants & products
Set the initial concentrations of all the chemicals, in their index order¶
dynamics = ReactionDynamics(reaction_data=chem_data)
dynamics.set_conc([50., 0.], snapshot=True)
dynamics.describe_state()
SYSTEM STATE at Time t = 0: 2 species: Species 0 (A). Conc: 50.0 Species 1 (B). Conc: 0.0
Run the reaction¶
dynamics.set_diagnostics() # To save diagnostic information about the call to single_compartment_react()
# All of these settings are currently close to the default values... but subject to change; set for repeatability
dynamics.set_thresholds(norm="norm_A", low=0.5, high=1.0, abort=1.44)
dynamics.set_thresholds(norm="norm_B", low=0.08, high=0.5, abort=1.5)
dynamics.set_step_factors(upshift=1.5, downshift=0.5, abort=0.5)
dynamics.set_error_step_factor(0.5)
# The changes of concentrations vary very rapidly early on; automated variable timesteps will take care of that
dynamics.single_compartment_react(initial_step=0.001, reaction_duration=0.3,
snapshots={"initial_caption": "1st reaction step",
"final_caption": "last reaction step"},
variable_steps=True, explain_variable_steps=False)
49 total step(s) taken
dynamics.get_history()
| SYSTEM TIME | A | B | caption | |
|---|---|---|---|---|
| 0 | 0.000000 | 50.000000 | 0.000000 | Initial state |
| 1 | 0.001000 | 48.500000 | 1.500000 | 1st reaction step |
| 2 | 0.001500 | 47.776250 | 2.223750 | |
| 3 | 0.002000 | 47.065166 | 2.934834 | |
| 4 | 0.002500 | 46.366525 | 3.633475 | |
| 5 | 0.003000 | 45.680111 | 4.319889 | |
| 6 | 0.003500 | 45.005709 | 4.994291 | |
| 7 | 0.004000 | 44.343109 | 5.656891 | |
| 8 | 0.004500 | 43.692105 | 6.307895 | |
| 9 | 0.005000 | 43.052493 | 6.947507 | |
| 10 | 0.005500 | 42.424074 | 7.575926 | |
| 11 | 0.006000 | 41.806653 | 8.193347 | |
| 12 | 0.006500 | 41.200037 | 8.799963 | |
| 13 | 0.007250 | 40.306036 | 9.693964 | |
| 14 | 0.008000 | 39.435502 | 10.564498 | |
| 15 | 0.008750 | 38.587820 | 11.412180 | |
| 16 | 0.009500 | 37.762390 | 12.237610 | |
| 17 | 0.010625 | 36.556746 | 13.443254 | |
| 18 | 0.011750 | 35.398574 | 14.601426 | |
| 19 | 0.012875 | 34.286005 | 15.713995 | |
| 20 | 0.014000 | 33.217244 | 16.782756 | |
| 21 | 0.015125 | 32.190565 | 17.809435 | |
| 22 | 0.016250 | 31.204311 | 18.795689 | |
| 23 | 0.017938 | 29.783182 | 20.216818 | |
| 24 | 0.018781 | 29.114585 | 20.885415 | |
| 25 | 0.020047 | 28.141306 | 21.858694 | |
| 26 | 0.021945 | 26.746057 | 23.253943 | |
| 27 | 0.023844 | 25.443516 | 24.556484 | |
| 28 | 0.025742 | 24.227523 | 25.772477 | |
| 29 | 0.027641 | 23.092327 | 26.907673 | |
| 30 | 0.029539 | 22.032560 | 27.967440 | |
| 31 | 0.031438 | 21.043209 | 28.956791 | |
| 32 | 0.034285 | 19.657789 | 30.342211 | |
| 33 | 0.037133 | 18.410451 | 31.589549 | |
| 34 | 0.039980 | 17.287433 | 32.712567 | |
| 35 | 0.042828 | 16.276344 | 33.723656 | |
| 36 | 0.045676 | 15.366028 | 34.633972 | |
| 37 | 0.049947 | 14.136648 | 35.863352 | |
| 38 | 0.054219 | 13.091062 | 36.908938 | |
| 39 | 0.058490 | 12.201794 | 37.798206 | |
| 40 | 0.064897 | 11.067313 | 38.932687 | |
| 41 | 0.071305 | 10.187242 | 39.812758 | |
| 42 | 0.080916 | 9.163174 | 40.836826 | |
| 43 | 0.090526 | 8.483581 | 41.516419 | |
| 44 | 0.104943 | 7.807093 | 42.192907 | |
| 45 | 0.126567 | 7.304364 | 42.695636 | |
| 46 | 0.159004 | 7.121008 | 42.878992 | |
| 47 | 0.207658 | 7.158215 | 42.841785 | |
| 48 | 0.280641 | 7.118985 | 42.881015 | |
| 49 | 0.390114 | 7.210452 | 42.789548 | last reaction step |
dynamics.explain_time_advance()
From time 0 to 0.001, in 1 step of 0.001 From time 0.001 to 0.0065, in 11 steps of 0.0005 From time 0.0065 to 0.0095, in 4 steps of 0.00075 From time 0.0095 to 0.01625, in 6 steps of 0.00113 From time 0.01625 to 0.01794, in 1 step of 0.00169 From time 0.01794 to 0.01878, in 1 step of 0.000844 From time 0.01878 to 0.02005, in 1 step of 0.00127 From time 0.02005 to 0.03144, in 6 steps of 0.0019 From time 0.03144 to 0.04568, in 5 steps of 0.00285 From time 0.04568 to 0.05849, in 3 steps of 0.00427 From time 0.05849 to 0.0713, in 2 steps of 0.00641 From time 0.0713 to 0.09053, in 2 steps of 0.00961 From time 0.09053 to 0.1049, in 1 step of 0.0144 From time 0.1049 to 0.1266, in 1 step of 0.0216 From time 0.1266 to 0.159, in 1 step of 0.0324 From time 0.159 to 0.2077, in 1 step of 0.0487 From time 0.2077 to 0.2806, in 1 step of 0.073 From time 0.2806 to 0.3901, in 1 step of 0.109 (49 steps total)
dynamics.plot_curves(colors=["blue", "green"], title="Single reaction A <-> B (no downregulation)",
show_intervals=True)
Notice the intersection at the exact midpoint of the 2 initial concentrations (50 and 0):¶
dynamics.curve_intersection('A', 'B', t_start=0, t_end=0.1)
Min abs distance found at data row: 27
(0.02453617850732098, 25.000000000000007)
# Verify that all the reactions have reached equilibrium
dynamics.is_in_equilibrium(tolerance=2)
A <-> B
Final concentrations: [B] = 42.79 ; [A] = 7.21
1. Ratio of reactant/product concentrations, adjusted for reaction orders: 5.93438
Formula used: [B] / [A]
2. Ratio of forward/reverse reaction rates: 6.0
Discrepancy between the two values: 1.094 %
Reaction IS in equilibrium (within 2% tolerance)
True
# Register the new chemical ("S")
chem_data.add_chemical("S")
# Add the reaction A <-> S (fast shunt, poor thermodynical energetic advantage)
chem_data.add_reaction(reactants=["A"], products=["S"],
forward_rate=150., reverse_rate=100.)
chem_data.describe_reactions()
Number of reactions: 2 (at temp. 25 C) 0: A <-> B (kF = 30 / kR = 5 / Delta_G = -4,441.69 / K = 6) | 1st order in all reactants & products 1: A <-> S (kF = 150 / kR = 100 / Delta_G = -1,005.13 / K = 1.5) | 1st order in all reactants & products
# Send a plot of the network of reactions to the HTML log file
graph_data = chem_data.prepare_graph_network()
GraphicLog.export_plot(graph_data, "vue_cytoscape_1")
[GRAPHIC ELEMENT SENT TO LOG FILE `down_regulate_1.log.htm`]
Set the initial concentrations of all the chemicals, in their index order¶
dynamics = ReactionDynamics(reaction_data=chem_data) # Notice we're over-writing the earlier "dynamics" object
dynamics.set_conc([50., 0, 0.], snapshot=True)
dynamics.describe_state()
SYSTEM STATE at Time t = 0: 3 species: Species 0 (A). Conc: 50.0 Species 1 (B). Conc: 0.0 Species 2 (S). Conc: 0.0
Run the reaction¶
dynamics.set_diagnostics() # To save diagnostic information about the call to single_compartment_react()
# All of these settings are currently close to the default values... but subject to change; set for repeatability
dynamics.set_thresholds(norm="norm_A", low=0.5, high=1.0, abort=1.44)
dynamics.set_thresholds(norm="norm_B", low=0.05, high=0.5, abort=1.5)
dynamics.set_step_factors(upshift=1.4, downshift=0.5, abort=0.5)
dynamics.set_error_step_factor(0.333)
# The changes of concentrations vary very rapidly early on; automated variable timesteps will take care of that
dynamics.single_compartment_react(initial_step=0.001, reaction_duration=0.3,
snapshots={"initial_caption": "1st reaction step",
"final_caption": "last reaction step"},
variable_steps=True, explain_variable_steps=False)
INFO: the tentative time step (0.001) leads to a least one norm value > its ABORT threshold:
-> will backtrack, and re-do step with a SMALLER delta time, multiplied by 0.5 (set to 0.0005) [Step started at t=0, and will rewind there]
INFO: the tentative time step (0.0005) leads to a least one norm value > its ABORT threshold:
-> will backtrack, and re-do step with a SMALLER delta time, multiplied by 0.5 (set to 0.00025) [Step started at t=0, and will rewind there]
69 total step(s) taken
dynamics.plot_curves(colors=["blue", "green", "red"],
title="Coupled reactions A <-> B and A <-> S (fast but disadvantaged energetically)",
show_intervals=True)
dynamics.explain_time_advance()
From time 0 to 0.0005, in 2 steps of 0.00025 From time 0.0005 to 0.00225, in 14 steps of 0.000125 From time 0.00225 to 0.00295, in 4 steps of 0.000175 From time 0.00295 to 0.00393, in 4 steps of 0.000245 From time 0.00393 to 0.005302, in 4 steps of 0.000343 From time 0.005302 to 0.006743, in 3 steps of 0.00048 From time 0.006743 to 0.009432, in 4 steps of 0.000672 From time 0.009432 to 0.01226, in 3 steps of 0.000941 From time 0.01226 to 0.01753, in 4 steps of 0.00132 From time 0.01753 to 0.0249, in 4 steps of 0.00184 From time 0.0249 to 0.03524, in 4 steps of 0.00258 From time 0.03524 to 0.04608, in 3 steps of 0.00362 From time 0.04608 to 0.06127, in 3 steps of 0.00506 From time 0.06127 to 0.07544, in 2 steps of 0.00709 From time 0.07544 to 0.1151, in 4 steps of 0.00992 From time 0.1151 to 0.1429, in 2 steps of 0.0139 From time 0.1429 to 0.1818, in 2 steps of 0.0194 From time 0.1818 to 0.209, in 1 step of 0.0272 From time 0.209 to 0.2471, in 1 step of 0.0381 From time 0.2471 to 0.3005, in 1 step of 0.0534 (69 steps total)
dynamics.get_history()
| SYSTEM TIME | A | B | S | caption | |
|---|---|---|---|---|---|
| 0 | 0.000000 | 50.000000 | 0.000000 | 0.000000 | Initial state |
| 1 | 0.000250 | 47.750000 | 0.375000 | 1.875000 | 1st reaction step |
| 2 | 0.000500 | 45.648594 | 0.732656 | 3.618750 | |
| 3 | 0.000625 | 44.667193 | 0.903381 | 4.429427 | |
| 4 | 0.000750 | 43.718113 | 1.070318 | 5.211569 | |
| ... | ... | ... | ... | ... | ... |
| 65 | 0.162354 | 6.593940 | 33.315058 | 10.091002 | |
| 66 | 0.181800 | 6.375490 | 33.922608 | 9.701902 | |
| 67 | 0.209024 | 6.163548 | 34.512062 | 9.324390 | |
| 68 | 0.247138 | 5.994349 | 34.982624 | 9.023027 | |
| 69 | 0.300498 | 5.900061 | 35.245013 | 8.854926 | last reaction step |
70 rows × 5 columns
# Verify that all the reactions have reached equilibrium
dynamics.is_in_equilibrium()
A <-> B
Final concentrations: [B] = 35.25 ; [A] = 5.9
1. Ratio of reactant/product concentrations, adjusted for reaction orders: 5.97367
Formula used: [B] / [A]
2. Ratio of forward/reverse reaction rates: 6.0
Discrepancy between the two values: 0.4388 %
Reaction IS in equilibrium (within 1% tolerance)
A <-> S
Final concentrations: [S] = 8.855 ; [A] = 5.9
1. Ratio of reactant/product concentrations, adjusted for reaction orders: 1.50082
Formula used: [S] / [A]
2. Ratio of forward/reverse reaction rates: 1.5
Discrepancy between the two values: 0.05463 %
Reaction IS in equilibrium (within 1% tolerance)
True
# Specify the chemicals (notice that we're starting with new objects)
chem_data = chem(names=["A", "B", "S"])
# Reaction A <-> B (as before)
chem_data.add_reaction(reactants=["A"], products=["B"],
forward_rate=30., reverse_rate=5.)
# Reaction A <-> S (slow shunt, excellent thermodynamical energetic advantage)
chem_data.add_reaction(reactants=["A"], products=["S"],
forward_rate=3., reverse_rate=0.1)
chem_data.describe_reactions()
Number of reactions: 2 (at temp. 25 C) 0: A <-> B (kF = 30 / kR = 5 / Delta_G = -4,441.69 / K = 6) | 1st order in all reactants & products 1: A <-> S (kF = 3 / kR = 0.1 / Delta_G = -8,431.42 / K = 30) | 1st order in all reactants & products
Set the initial concentrations of all the chemicals, in their index order¶
dynamics = ReactionDynamics(reaction_data=chem_data)
dynamics.set_conc([50., 0, 0.], snapshot=True)
dynamics.describe_state()
SYSTEM STATE at Time t = 0: 3 species: Species 0 (A). Conc: 50.0 Species 1 (B). Conc: 0.0 Species 2 (S). Conc: 0.0
Run the reaction¶
dynamics.set_diagnostics() # To save diagnostic information about the call to single_compartment_react()
# These settings can be tweaked to make the time resolution finer or coarser
dynamics.set_thresholds(norm="norm_A", low=2.0, high=5.0, abort=10.0)
dynamics.set_thresholds(norm="norm_B", low=0.008, high=0.5, abort=2.0) # The "low" value here seems especially critical to fend off instabilities
dynamics.set_step_factors(upshift=1.5, downshift=0.25, abort=0.25)
dynamics.set_error_step_factor(0.2)
# The changes of concentrations vary very rapidly early on; automated variable timesteps will take care of that
dynamics.single_compartment_react(initial_step=0.005, reaction_duration=7.0,
snapshots={"initial_caption": "1st reaction step",
"final_caption": "last reaction step"},
variable_steps=True, explain_variable_steps=False)
INFO: the tentative time step (0.005) leads to a least one norm value > its ABORT threshold:
-> will backtrack, and re-do step with a SMALLER delta time, multiplied by 0.25 (set to 0.00125) [Step started at t=0, and will rewind there]
454 total step(s) taken
dynamics.plot_curves(colors=["blue", "green", "red"],
title="Coupled reactions A <-> B and A <-> S (slow but with energetic advantage)")
dynamics.explain_time_advance()
From time 0 to 0.0025, in 2 steps of 0.00125 From time 0.0025 to 0.03281, in 97 steps of 0.000313 From time 0.03281 to 0.05859, in 55 steps of 0.000469 From time 0.05859 to 0.07617, in 25 steps of 0.000703 From time 0.07617 to 0.09199, in 15 steps of 0.00105 From time 0.09199 to 0.1062, in 9 steps of 0.00158 From time 0.1062 to 0.1205, in 6 steps of 0.00237 From time 0.1205 to 0.2237, in 29 steps of 0.00356 From time 0.2237 to 0.3999, in 33 steps of 0.00534 From time 0.3999 to 0.6402, in 30 steps of 0.00801 From time 0.6402 to 0.9525, in 26 steps of 0.012 From time 0.9525 to 1.349, in 22 steps of 0.018 From time 1.349 to 1.998, in 24 steps of 0.027 From time 1.998 to 3.66, in 41 steps of 0.0405 From time 3.66 to 4.876, in 20 steps of 0.0608 From time 4.876 to 5.971, in 12 steps of 0.0912 From time 5.971 to 6.929, in 7 steps of 0.137 From time 6.929 to 7.134, in 1 step of 0.205 (454 steps total)
dynamics.get_history()
| SYSTEM TIME | A | B | S | caption | |
|---|---|---|---|---|---|
| 0 | 0.000000 | 50.000000 | 0.000000 | 0.000000 | Initial state |
| 1 | 0.001250 | 47.937500 | 1.875000 | 0.187500 | 1st reaction step |
| 2 | 0.002500 | 45.971820 | 3.660938 | 0.367242 | |
| 3 | 0.002812 | 45.503468 | 4.086203 | 0.410329 | |
| 4 | 0.003125 | 45.040611 | 4.506413 | 0.452976 | |
| ... | ... | ... | ... | ... | ... |
| 450 | 6.518546 | 1.538995 | 9.356824 | 39.104182 | |
| 451 | 6.655388 | 1.526366 | 9.272764 | 39.200870 | |
| 452 | 6.792230 | 1.514574 | 9.194376 | 39.291050 | |
| 453 | 6.929072 | 1.503631 | 9.121214 | 39.375155 | |
| 454 | 7.134334 | 1.487983 | 9.019168 | 39.492848 | last reaction step |
455 rows × 5 columns
Check the final equilibrium¶
# Verify that all the reactions have reached equilibrium
dynamics.is_in_equilibrium(tolerance=12)
A <-> B
Final concentrations: [B] = 9.019 ; [A] = 1.488
1. Ratio of reactant/product concentrations, adjusted for reaction orders: 6.06134
Formula used: [B] / [A]
2. Ratio of forward/reverse reaction rates: 6.0
Discrepancy between the two values: 1.022 %
Reaction IS in equilibrium (within 12% tolerance)
A <-> S
Final concentrations: [S] = 39.49 ; [A] = 1.488
1. Ratio of reactant/product concentrations, adjusted for reaction orders: 26.5412
Formula used: [S] / [A]
2. Ratio of forward/reverse reaction rates: 30.0
Discrepancy between the two values: 11.53 %
Reaction IS in equilibrium (within 12% tolerance)
True
Please note the much-longer timescale from the earlier plots¶
If we look at early time interval, this is what it looks like:
fig = px.line(data_frame=dynamics.get_history().loc[:250], x="SYSTEM TIME", y=["A", "B", "S"],
title="Same plot as above, both only showing initial detail",
color_discrete_sequence = ["blue", "green", "red"],
labels={"value":"concentration", "variable":"Chemical"})
fig.show()
# Look at where the curves intersect
dynamics.curve_intersection("A", "B", t_start=0, t_end=0.1)
Min abs distance found at data row: 71
(0.02405919499545674, 23.73396682504195)
dynamics.curve_intersection("A", "S", t_start=0.1, t_end=0.2)
Min abs distance found at data row: 216
(0.14412951669101942, 6.026379520544665)