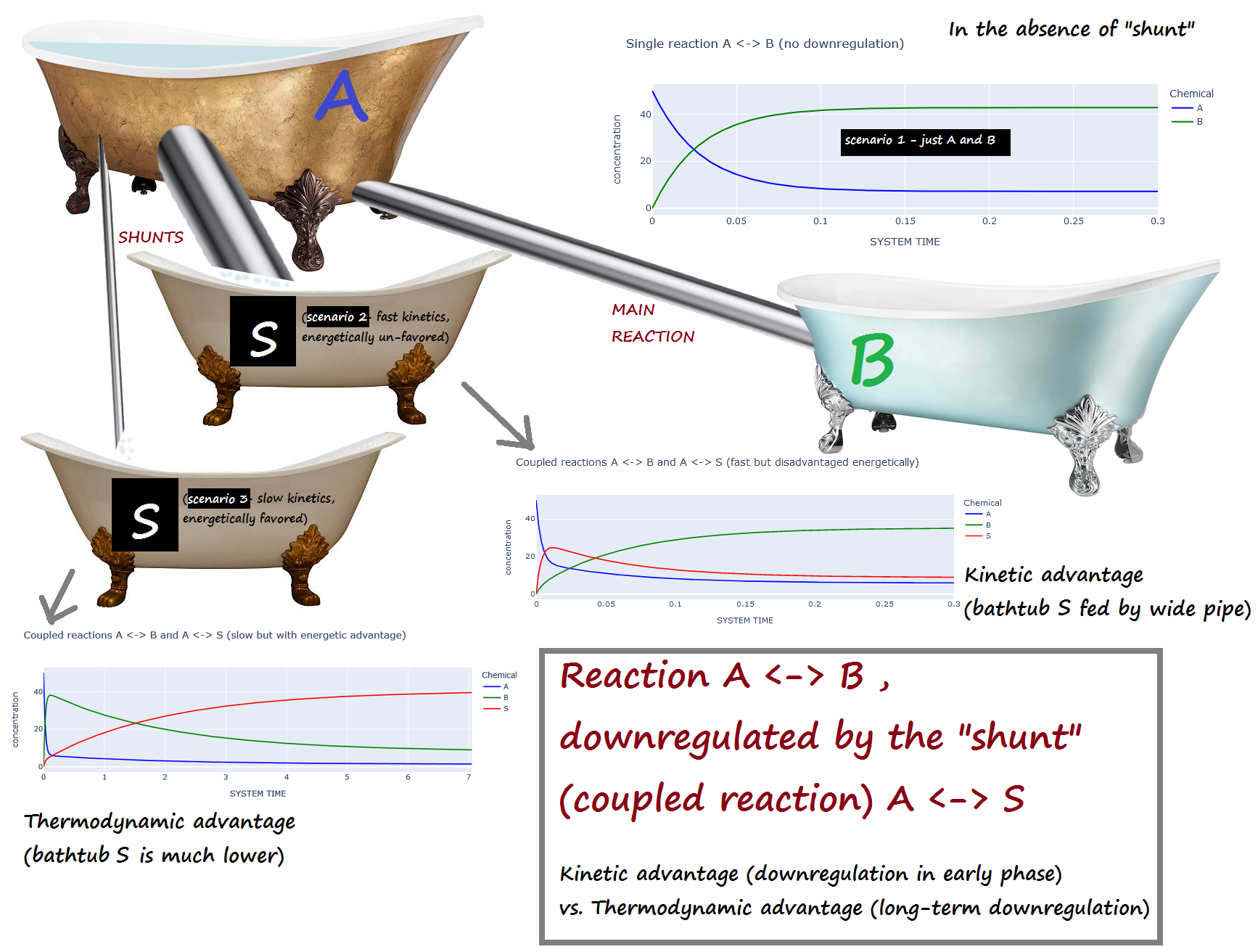
A <-> B , downregulated by the "shunt" (coupled reaction) A <-> S¶
Kinetic advantage (downregulation in early phase) vs. Thermodynamic advantage (long-term downregulation)¶
Scenario 1 : No downregulation on A <-> B
Scenario 2 : The shunt (A <-> S) has a kinetic advantage but thermodynamic DIS-advantage compared to A <-> B
(i.e. A <-> S is fast, but energetically unfavored)
Scenario 3 : The shunt (A <-> S) is has a kinetic DIS-advantage but a thermodynamic advantage compared to A <-> B
(i.e. A <-> S is slow, but energetically favored)
All reactions 1st order, mostly forward. Taken to equilibrium.
LAST REVISED: Feb. 5, 2023
Bathtub analogy:¶
A is initially full, while B and S are empty.
If the "shunt" S is present, scenario 2 corresponds to a large pipe and a small elevation change...
while scenario 3 corresponds to a narrow pipe and a large elevation change.
# Extend the sys.path variable, to contain the project's root directory
import set_path
set_path.add_ancestor_dir_to_syspath(2) # The number of levels to go up
# to reach the project's home, from the folder containing this notebook
Added 'D:\Docs\- MY CODE\BioSimulations\life123-Win7' to sys.path
from experiments.get_notebook_info import get_notebook_basename
from src.modules.reactions.reaction_data import ReactionData as chem
from src.modules.reactions.reaction_dynamics import ReactionDynamics
import numpy as np
import plotly.express as px
from src.modules.visualization.graphic_log import GraphicLog
# Initialize the HTML logging (for the graphics)
log_file = get_notebook_basename() + ".log.htm" # Use the notebook base filename for the log file
# Set up the use of some specified graphic (Vue) components
GraphicLog.config(filename=log_file,
components=["vue_cytoscape_1"],
extra_js="https://cdnjs.cloudflare.com/ajax/libs/cytoscape/3.21.2/cytoscape.umd.js")
-> Output will be LOGGED into the file 'down_regulate_1.log.htm'
Scenario 1: A <-> B in the absence of the 2nd reaction¶
Initialize the System¶
Specify the chemicals and the reaction
# Specify the chemicals
chem_data = chem(names=["A", "B"])
# Reaction A <-> B
chem_data.add_reaction(reactants=["A"], products=["B"],
forward_rate=30., reverse_rate=5.)
chem_data.describe_reactions()
Number of reactions: 1 (at temp. 25 C) 0: A <-> B (kF = 30 / kR = 5 / Delta_G = -4,441.69 / K = 6) | 1st order in all reactants & products
Set the initial concentrations of all the chemicals, in their index order¶
dynamics = ReactionDynamics(reaction_data=chem_data)
dynamics.set_conc([50., 0.], snapshot=True)
dynamics.describe_state()
SYSTEM STATE at Time t = 0: 2 species: Species 0 (A). Conc: 50.0 Species 1 (B). Conc: 0.0
Run the reaction¶
dynamics.set_diagnostics() # To save diagnostic information about the call to single_compartment_react()
#dynamics.verbose_list = [1, 2, 3] # Uncomment for detailed run information (meant for debugging the adaptive variable time step)
# The changes of concentrations vary very rapidly early on;
# so, we'll be using the dynamic_substeps option to increase time resolution,
# as long as the reaction remains "fast" (based on a threshold of % change, as specified by fast_threshold)
dynamics.single_compartment_react(time_step=0.001, reaction_duration=0.05,
snapshots={"initial_caption": "1st reaction step",
"final_caption": "last reaction step"},
dynamic_substeps=4, rel_fast_threshold=25)
single_compartment_react(): setting abs_fast_threshold to 250.0 50 total step(s) taken
df_iterm = dynamics.get_history()
df_iterm
| SYSTEM TIME | A | B | caption | |
|---|---|---|---|---|
| 0 | 0.00000 | 50.000000 | 0.000000 | Initial state |
| 1 | 0.00025 | 49.625000 | 0.375000 | Interm. step, due to the fast rxns: [0] |
| 2 | 0.00050 | 49.253281 | 0.746719 | Interm. step, due to the fast rxns: [0] |
| 3 | 0.00075 | 48.884815 | 1.115185 | Interm. step, due to the fast rxns: [0] |
| 4 | 0.00100 | 48.519573 | 1.480427 | 1st reaction step |
| ... | ... | ... | ... | ... |
| 196 | 0.04900 | 14.797598 | 35.202402 | |
| 197 | 0.04925 | 14.730619 | 35.269381 | Interm. step, due to the fast rxns: [0] |
| 198 | 0.04950 | 14.664226 | 35.335774 | Interm. step, due to the fast rxns: [0] |
| 199 | 0.04975 | 14.598414 | 35.401586 | Interm. step, due to the fast rxns: [0] |
| 200 | 0.05000 | 14.533178 | 35.466822 | last reaction step |
201 rows × 4 columns
# Continue running the reaction at lover resolution
dynamics.single_compartment_react(time_step=0.002, reaction_duration=0.25,
snapshots={"initial_caption": "1st reaction step",
"final_caption": "last reaction step"},
dynamic_substeps=4, rel_fast_threshold=10)
single_compartment_react(): setting abs_fast_threshold to 50.0 125 total step(s) taken
df = dynamics.get_history()
df
| SYSTEM TIME | A | B | caption | |
|---|---|---|---|---|
| 0 | 0.00000 | 50.000000 | 0.000000 | Initial state |
| 1 | 0.00025 | 49.625000 | 0.375000 | Interm. step, due to the fast rxns: [0] |
| 2 | 0.00050 | 49.253281 | 0.746719 | Interm. step, due to the fast rxns: [0] |
| 3 | 0.00075 | 48.884815 | 1.115185 | Interm. step, due to the fast rxns: [0] |
| 4 | 0.00100 | 48.519573 | 1.480427 | 1st reaction step |
| ... | ... | ... | ... | ... |
| 393 | 0.29200 | 7.144047 | 42.855953 | |
| 394 | 0.29400 | 7.143964 | 42.856036 | |
| 395 | 0.29600 | 7.143886 | 42.856114 | |
| 396 | 0.29800 | 7.143814 | 42.856186 | |
| 397 | 0.30000 | 7.143747 | 42.856253 | last reaction step |
398 rows × 4 columns
dynamics.explain_time_advance()
From time 0 to 0.05, in 200 substeps of 0.00025 (each 1/4 of full step) From time 0.05 to 0.098, in 96 substeps of 0.0005 (each 1/4 of full step) From time 0.098 to 0.3, in 101 FULL steps of 0.002
# Verify that all the reactions have reached equilibrium
dynamics.is_in_equilibrium()
A <-> B
Final concentrations: [B] = 42.86 ; [A] = 7.144
1. Ratio of reactant/product concentrations, adjusted for reaction orders: 5.99913
Formula used: [B] / [A]
2. Ratio of forward/reverse reaction rates: 6.0
Discrepancy between the two values: 0.01453 %
Reaction IS in equilibrium (within 1% tolerance)
True
dynamics.plot_curves(colors=["blue", "green"], title="Single reaction A <-> B (no downregulation)")
# Register the new chemical ("S")
chem_data.add_chemical("S")
# Add the reaction A <-> S (fast shunt, poor thermodynical energetic advantage)
chem_data.add_reaction(reactants=["A"], products=["S"],
forward_rate=150., reverse_rate=100.)
chem_data.describe_reactions()
Number of reactions: 2 (at temp. 25 C) 0: A <-> B (kF = 30 / kR = 5 / Delta_G = -4,441.69 / K = 6) | 1st order in all reactants & products 1: A <-> S (kF = 150 / kR = 100 / Delta_G = -1,005.13 / K = 1.5) | 1st order in all reactants & products
# Send a plot of the network of reactions to the HTML log file
graph_data = chem_data.prepare_graph_network()
GraphicLog.export_plot(graph_data, "vue_cytoscape_1")
[GRAPHIC ELEMENT SENT TO LOG FILE `down_regulate_1.log.htm`]
Set the initial concentrations of all the chemicals, in their index order¶
dynamics = ReactionDynamics(reaction_data=chem_data) # Notice we're over-writing the earlier "dynamics" object
dynamics.set_conc([50., 0, 0.], snapshot=True)
dynamics.describe_state()
SYSTEM STATE at Time t = 0: 3 species: Species 0 (A). Conc: 50.0 Species 1 (B). Conc: 0.0 Species 2 (S). Conc: 0.0
Run the reaction¶
dynamics.set_diagnostics() # To save diagnostic information about the call to single_compartment_react()
#dynamics.verbose_list = [1, 2, 3] # Uncomment for detailed run information (meant for debugging the adaptive variable time step)
# The changes of concentrations vary very rapidly early on;
# so, we'll be using the dynamic_substeps option to increase time resolution,
# as long as the reaction remains "fast" (based on a threshold of % change, as specified by fast_threshold)
dynamics.single_compartment_react(time_step=0.001, reaction_duration=0.05,
snapshots={"initial_caption": "1st reaction step",
"final_caption": "last reaction step"},
dynamic_substeps=4, rel_fast_threshold=10)
single_compartment_react(): setting abs_fast_threshold to 100.0 50 total step(s) taken
# Continue running the reaction at lover resolution
dynamics.single_compartment_react(time_step=0.002, reaction_duration=0.25,
snapshots={"initial_caption": "1st reaction step",
"final_caption": "last reaction step"},
dynamic_substeps=4, rel_fast_threshold=10)
single_compartment_react(): setting abs_fast_threshold to 50.0 125 total step(s) taken
df = dynamics.get_history()
df
| SYSTEM TIME | A | B | S | caption | |
|---|---|---|---|---|---|
| 0 | 0.00000 | 50.000000 | 0.000000 | 0.000000 | Initial state |
| 1 | 0.00025 | 47.750000 | 0.375000 | 1.875000 | Interm. step, due to the fast rxns: [0, 1] |
| 2 | 0.00050 | 45.648594 | 0.732656 | 3.618750 | Interm. step, due to the fast rxns: [0, 1] |
| 3 | 0.00075 | 43.685792 | 1.074105 | 5.240104 | Interm. step, due to the fast rxns: [0, 1] |
| 4 | 0.00100 | 41.852276 | 1.400406 | 6.747318 | 1st reaction step |
| ... | ... | ... | ... | ... | ... |
| 465 | 0.29200 | 5.990345 | 34.993770 | 9.015885 | |
| 466 | 0.29400 | 5.986936 | 35.003253 | 9.009812 | |
| 467 | 0.29600 | 5.983634 | 35.012436 | 9.003930 | |
| 468 | 0.29800 | 5.980436 | 35.021330 | 8.998234 | |
| 469 | 0.30000 | 5.977339 | 35.029943 | 8.992718 | last reaction step |
470 rows × 5 columns
Check the final equilibrium¶
# Verify that all the reactions have reached equilibrium
dynamics.is_in_equilibrium(tolerance=3)
A <-> B
Final concentrations: [B] = 35.03 ; [A] = 5.977
1. Ratio of reactant/product concentrations, adjusted for reaction orders: 5.86046
Formula used: [B] / [A]
2. Ratio of forward/reverse reaction rates: 6.0
Discrepancy between the two values: 2.326 %
Reaction IS in equilibrium (within 3% tolerance)
A <-> S
Final concentrations: [S] = 8.993 ; [A] = 5.977
1. Ratio of reactant/product concentrations, adjusted for reaction orders: 1.50447
Formula used: [S] / [A]
2. Ratio of forward/reverse reaction rates: 1.5
Discrepancy between the two values: 0.2979 %
Reaction IS in equilibrium (within 3% tolerance)
True
Plots of changes of concentration with time¶
dynamics.plot_curves(colors=["blue", "green", "red"],
title="Coupled reactions A <-> B and A <-> S (fast but disadvantaged energetically)")
# Verify that the stoichiometry is respected at every reaction step/substep (NOTE: it requires earlier activation of saving diagnostic data)
dynamics.stoichiometry_checker_entire_run()
True
# Specify the chemicals (notice that we're starting with new objects)
chem_data3 = chem(names=["A", "B", "S"])
# Reaction A <-> B (as before)
chem_data3.add_reaction(reactants=["A"], products=["B"],
forward_rate=30., reverse_rate=5.)
# Reaction A <-> S (slow shunt, excellent thermodynamical energetic advantage)
chem_data3.add_reaction(reactants=["A"], products=["S"],
forward_rate=3., reverse_rate=0.1)
chem_data3.describe_reactions()
Number of reactions: 2 (at temp. 25 C) 0: A <-> B (kF = 30 / kR = 5 / Delta_G = -4,441.69 / K = 6) | 1st order in all reactants & products 1: A <-> S (kF = 3 / kR = 0.1 / Delta_G = -8,431.42 / K = 30) | 1st order in all reactants & products
Set the initial concentrations of all the chemicals, in their index order¶
dynamics3 = ReactionDynamics(reaction_data=chem_data3)
dynamics3.set_conc([50., 0, 0.], snapshot=True)
dynamics3.describe_state()
SYSTEM STATE at Time t = 0: 3 species: Species 0 (A). Conc: 50.0 Species 1 (B). Conc: 0.0 Species 2 (S). Conc: 0.0
Run the reaction¶
dynamics3.set_diagnostics() # To save diagnostic information about the call to single_compartment_react()
#dynamics3.verbose_list = [1, 2, 3] # Uncomment for detailed run information (meant for debugging the adaptive variable time step)
# The changes of concentrations vary very rapidly early on;
# so, we'll be using the dynamic_substeps option to increase time resolution,
# as long as the reaction remains "fast" (based on a threshold of % change, as specified by fast_threshold)
dynamics3.single_compartment_react(time_step=0.005, reaction_duration=0.3,
snapshots={"initial_caption": "1st reaction step",
"final_caption": "last reaction step"},
dynamic_substeps=5, rel_fast_threshold=10)
single_compartment_react(): setting abs_fast_threshold to 20.0 60 total step(s) taken
# Continue running the reaction at lover resolution
dynamics3.single_compartment_react(time_step=0.25, reaction_duration=6.7,
snapshots={"initial_caption": "1st reaction step",
"final_caption": "last reaction step"},
dynamic_substeps=5, rel_fast_threshold=10)
single_compartment_react(): setting abs_fast_threshold to 0.4 27 total step(s) taken
df3 = dynamics3.get_history()
df3
| SYSTEM TIME | A | B | S | caption | |
|---|---|---|---|---|---|
| 0 | 0.000 | 50.000000 | 0.000000 | 0.000000 | Initial state |
| 1 | 0.001 | 48.350000 | 1.500000 | 0.150000 | Interm. step, due to the fast rxns: [0, 1] |
| 2 | 0.002 | 46.761965 | 2.943000 | 0.295035 | Interm. step, due to the fast rxns: [0, 1] |
| 3 | 0.003 | 45.233565 | 4.331144 | 0.435291 | Interm. step, due to the fast rxns: [0, 1] |
| 4 | 0.004 | 43.762556 | 5.666495 | 0.570949 | Interm. step, due to the fast rxns: [0, 1] |
| ... | ... | ... | ... | ... | ... |
| 275 | 6.850 | 1.511216 | 9.171961 | 39.316824 | Interm. step, due to the fast rxns: [0, 1] |
| 276 | 6.900 | 1.507284 | 9.145794 | 39.346922 | Interm. step, due to the fast rxns: [0, 1] |
| 277 | 6.950 | 1.503449 | 9.120272 | 39.376280 | Interm. step, due to the fast rxns: [0, 1] |
| 278 | 7.000 | 1.499708 | 9.095377 | 39.404916 | Interm. step, due to the fast rxns: [0, 1] |
| 279 | 7.050 | 1.496059 | 9.071094 | 39.432847 | last reaction step |
280 rows × 5 columns
Check the final equilibrium¶
# Verify that all the reactions have reached equilibrium
dynamics3.is_in_equilibrium(tolerance=13)
A <-> B
Final concentrations: [B] = 9.071 ; [A] = 1.496
1. Ratio of reactant/product concentrations, adjusted for reaction orders: 6.06333
Formula used: [B] / [A]
2. Ratio of forward/reverse reaction rates: 6.0
Discrepancy between the two values: 1.055 %
Reaction IS in equilibrium (within 13% tolerance)
A <-> S
Final concentrations: [S] = 39.43 ; [A] = 1.496
1. Ratio of reactant/product concentrations, adjusted for reaction orders: 26.3578
Formula used: [S] / [A]
2. Ratio of forward/reverse reaction rates: 30.0
Discrepancy between the two values: 12.14 %
Reaction IS in equilibrium (within 13% tolerance)
True
Plots of changes of concentration with time¶
dynamics3.plot_curves(colors=["blue", "green", "red"],
title="Coupled reactions A <-> B and A <-> S (slow but with energetic advantage)")
Please note the much-longer timescale from the earlier plots¶
If we look at the initial [0-0.1] interval, this is what it looks like:
fig = px.line(data_frame=dynamics3.get_history().loc[:96], x="SYSTEM TIME", y=["A", "B", "S"],
title="Same as above, both only showing initial detail",
color_discrete_sequence = ['blue', 'green', 'red'],
labels={"value":"concentration", "variable":"Chemical"})
fig.show()
dynamics3.curve_intersection(t_start=0, t_end=0.08, var1="A", var2="B")
Min abs distance found at row: 24
(0.02378914647211626, 23.73644141393734)
# Verify that the stoichiometry is respected at every reaction step/substep (NOTE: it requires earlier activation of saving diagnostic data)
dynamics.stoichiometry_checker_entire_run()
True