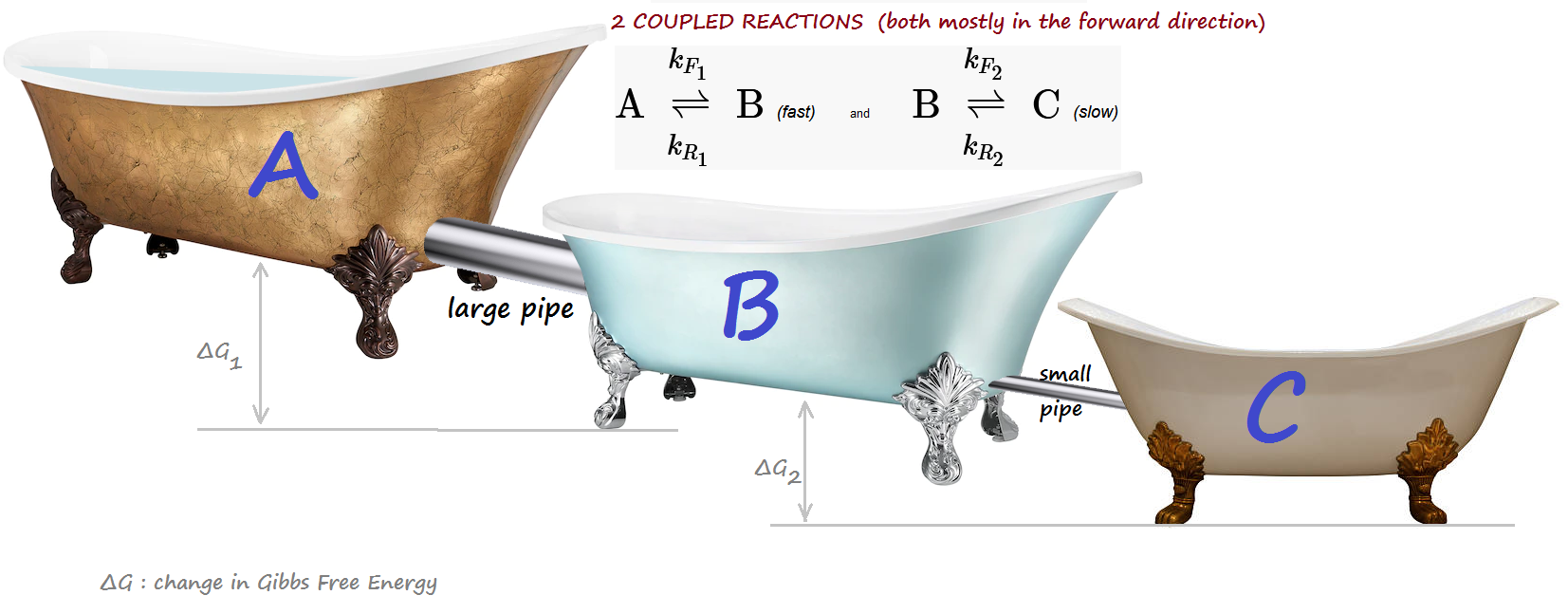
2 COUPLED reactions of different speeds, forming a "cascade":¶
A <-> B (fast) and B <-> C (slow)¶
All 1st order. Taken to equilibrium. Both reactions are mostly forward. The concentration of the intermediate product B manifests 1 oscillation (transient "overshoot")
(Adaptive variable time resolution is used, with extensive diagnostics.)
LAST REVISED: Feb. 5, 2023
Bathtub analogy:¶
A is initially full, while B and C are empty.
Tubs are progressively lower (reactions are mostly forward.)
A BIG pipe connects A and B: fast kinetics. A small pipe connects B and C: slow kinetics.
INTUITION: B, unable to quickly drain into C while at the same time being blasted by a hefty inflow from A,
will experience a transient surge, in excess of its final equilibrium level.
# Extend the sys.path variable, to contain the project's root directory
import set_path
set_path.add_ancestor_dir_to_syspath(2) # The number of levels to go up
# to reach the project's home, from the folder containing this notebook
Added 'D:\Docs\- MY CODE\BioSimulations\life123-Win7' to sys.path
from experiments.get_notebook_info import get_notebook_basename
from src.modules.reactions.reaction_data import ReactionData as chem
from src.modules.reactions.reaction_dynamics import ReactionDynamics
import numpy as np
import plotly.express as px
from src.modules.visualization.graphic_log import GraphicLog
# Initialize the HTML logging (for the graphics)
log_file = get_notebook_basename() + ".log.htm" # Use the notebook base filename for the log file
# Set up the use of some specified graphic (Vue) components
GraphicLog.config(filename=log_file,
components=["vue_cytoscape_1"],
extra_js="https://cdnjs.cloudflare.com/ajax/libs/cytoscape/3.21.2/cytoscape.umd.js")
-> Output will be LOGGED into the file 'cascade_1.log.htm'
Initialize the System¶
Specify the chemicals and the reactions
# Specify the chemicals
chem_data = chem(names=["A", "B", "C"])
# Reaction A <-> B (fast)
chem_data.add_reaction(reactants=["A"], products=["B"],
forward_rate=64., reverse_rate=8.)
# Reaction B <-> C (slow)
chem_data.add_reaction(reactants=["B"], products=["C"],
forward_rate=12., reverse_rate=2.)
print("Number of reactions: ", chem_data.number_of_reactions())
Number of reactions: 2
chem_data.describe_reactions()
Number of reactions: 2 (at temp. 25 C) 0: A <-> B (kF = 64 / kR = 8 / Delta_G = -5,154.85 / K = 8) | 1st order in all reactants & products 1: B <-> C (kF = 12 / kR = 2 / Delta_G = -4,441.69 / K = 6) | 1st order in all reactants & products
# Send a plot of the network of reactions to the HTML log file
graph_data = chem_data.prepare_graph_network()
GraphicLog.export_plot(graph_data, "vue_cytoscape_1")
[GRAPHIC ELEMENT SENT TO LOG FILE `cascade_1.log.htm`]
Start the simulation¶
dynamics = ReactionDynamics(reaction_data=chem_data)
Set the initial concentrations of all the chemicals, in their index order¶
dynamics.set_conc([50., 0, 0.], snapshot=True)
dynamics.describe_state()
SYSTEM STATE at Time t = 0: 3 species: Species 0 (A). Conc: 50.0 Species 1 (B). Conc: 0.0 Species 2 (C). Conc: 0.0
dynamics.get_history()
| SYSTEM TIME | A | B | C | caption | |
|---|---|---|---|---|---|
| 0 | 0.0 | 50.0 | 0.0 | 0.0 | Initial state |
Run the reaction¶
dynamics.set_diagnostics() # To save diagnostic information about the call to single_compartment_react()
#dynamics.verbose_list = [1, 2, 3] # Uncomment for detailed run information (meant for debugging the adaptive variable time step)
# The changes of concentrations vary very rapidly early on;
# so, we'll be using the dynamic_substeps option to increase time resolution,
# as long as the reaction remains "fast" (based on a threshold of % change, as specified by fast_threshold)
dynamics.single_compartment_react(time_step=0.02, reaction_duration=0.4,
snapshots={"initial_caption": "1st reaction step",
"final_caption": "last reaction step"},
dynamic_substeps=10, rel_fast_threshold=190)
single_compartment_react(): setting abs_fast_threshold to 95.0 20 total step(s) taken
Note: the argument dynamic_step=10 splits the time steps in 10 for any reactions that are "fast-changing" (as determined using the given value of fast_threshold )¶
df = dynamics.get_history()
df
| SYSTEM TIME | A | B | C | caption | |
|---|---|---|---|---|---|
| 0 | 0.000 | 50.000000 | 0.000000 | 0.000000 | Initial state |
| 1 | 0.002 | 43.600000 | 6.400000 | 0.000000 | Interm. step, due to the fast rxns: [0, 1] |
| 2 | 0.004 | 38.121600 | 11.724800 | 0.153600 | Interm. step, due to the fast rxns: [0, 1] |
| 3 | 0.006 | 33.429632 | 16.135987 | 0.434381 | Interm. step, due to the fast rxns: [0, 1] |
| 4 | 0.008 | 29.408815 | 19.771278 | 0.819907 | Interm. step, due to the fast rxns: [0, 1] |
| ... | ... | ... | ... | ... | ... |
| 88 | 0.320 | 0.970191 | 7.617998 | 41.411811 | |
| 89 | 0.340 | 0.947226 | 7.469116 | 41.583658 | |
| 90 | 0.360 | 0.929835 | 7.357265 | 41.712899 | |
| 91 | 0.380 | 0.916809 | 7.273064 | 41.810127 | |
| 92 | 0.400 | 0.906984 | 7.209759 | 41.883257 | last reaction step |
93 rows × 5 columns
dynamics.explain_time_advance()
From time 0 to 0.16, in 80 substeps of 0.002 (each 1/10 of full step) From time 0.16 to 0.4, in 12 FULL steps of 0.02
# Expand the early part
df.loc[:59]
| SYSTEM TIME | A | B | C | caption | |
|---|---|---|---|---|---|
| 0 | 0.000 | 50.000000 | 0.000000 | 0.000000 | Initial state |
| 1 | 0.002 | 43.600000 | 6.400000 | 0.000000 | Interm. step, due to the fast rxns: [0, 1] |
| 2 | 0.004 | 38.121600 | 11.724800 | 0.153600 | Interm. step, due to the fast rxns: [0, 1] |
| 3 | 0.006 | 33.429632 | 16.135987 | 0.434381 | Interm. step, due to the fast rxns: [0, 1] |
| 4 | 0.008 | 29.408815 | 19.771278 | 0.819907 | Interm. step, due to the fast rxns: [0, 1] |
| 5 | 0.010 | 25.960827 | 22.748035 | 1.291138 | Interm. step, due to the fast rxns: [0, 1] |
| 6 | 0.012 | 23.001810 | 25.166264 | 1.831926 | Interm. step, due to the fast rxns: [0, 1] |
| 7 | 0.014 | 20.460238 | 27.111173 | 2.428589 | Interm. step, due to the fast rxns: [0, 1] |
| 8 | 0.016 | 18.275107 | 28.655351 | 3.069543 | Interm. step, due to the fast rxns: [0, 1] |
| 9 | 0.018 | 16.394379 | 29.860628 | 3.744993 | Interm. step, due to the fast rxns: [0, 1] |
| 10 | 0.020 | 14.773668 | 30.779664 | 4.446668 | 1st reaction step |
| 11 | 0.022 | 13.375113 | 31.457293 | 5.167593 | Interm. step, due to the fast rxns: [0, 1] |
| 12 | 0.024 | 12.166415 | 31.931687 | 5.901898 | Interm. step, due to the fast rxns: [0, 1] |
| 13 | 0.026 | 11.120021 | 32.235328 | 6.644651 | Interm. step, due to the fast rxns: [0, 1] |
| 14 | 0.028 | 10.212424 | 32.395856 | 7.391720 | Interm. step, due to the fast rxns: [0, 1] |
| 15 | 0.030 | 9.423567 | 32.436779 | 8.139654 | Interm. step, due to the fast rxns: [0, 1] |
| 16 | 0.032 | 8.736339 | 32.378083 | 8.885578 | Interm. step, due to the fast rxns: [0, 1] |
| 17 | 0.034 | 8.136137 | 32.236753 | 9.627110 | Interm. step, due to the fast rxns: [0, 1] |
| 18 | 0.036 | 7.610500 | 32.027217 | 10.362283 | Interm. step, due to the fast rxns: [0, 1] |
| 19 | 0.038 | 7.148791 | 31.761722 | 11.089487 | Interm. step, due to the fast rxns: [0, 1] |
| 20 | 0.040 | 6.741933 | 31.450656 | 11.807411 | |
| 21 | 0.042 | 6.382176 | 31.102827 | 12.514997 | Interm. step, due to the fast rxns: [0, 1] |
| 22 | 0.044 | 6.062903 | 30.725692 | 13.211405 | Interm. step, due to the fast rxns: [0, 1] |
| 23 | 0.046 | 5.778463 | 30.325562 | 13.895976 | Interm. step, due to the fast rxns: [0, 1] |
| 24 | 0.048 | 5.524028 | 29.907766 | 14.568205 | Interm. step, due to the fast rxns: [0, 1] |
| 25 | 0.050 | 5.295477 | 29.476804 | 15.227719 | Interm. step, due to the fast rxns: [0, 1] |
| 26 | 0.052 | 5.089285 | 29.036464 | 15.874251 | Interm. step, due to the fast rxns: [0, 1] |
| 27 | 0.054 | 4.902440 | 28.589931 | 16.507629 | Interm. step, due to the fast rxns: [0, 1] |
| 28 | 0.056 | 4.732366 | 28.139876 | 17.127757 | Interm. step, due to the fast rxns: [0, 1] |
| 29 | 0.058 | 4.576861 | 27.688535 | 17.734603 | Interm. step, due to the fast rxns: [0, 1] |
| 30 | 0.060 | 4.434040 | 27.237771 | 18.328190 | |
| 31 | 0.062 | 4.302287 | 26.789130 | 18.908583 | Interm. step, due to the fast rxns: [1] |
| 32 | 0.064 | 4.170534 | 26.353578 | 19.475888 | Interm. step, due to the fast rxns: [1] |
| 33 | 0.066 | 4.038781 | 25.930748 | 20.030470 | Interm. step, due to the fast rxns: [1] |
| 34 | 0.068 | 3.907029 | 25.520285 | 20.572687 | Interm. step, due to the fast rxns: [1] |
| 35 | 0.070 | 3.775276 | 25.121841 | 21.102883 | Interm. step, due to the fast rxns: [1] |
| 36 | 0.072 | 3.643523 | 24.735082 | 21.621395 | Interm. step, due to the fast rxns: [1] |
| 37 | 0.074 | 3.511770 | 24.359678 | 22.128552 | Interm. step, due to the fast rxns: [1] |
| 38 | 0.076 | 3.380018 | 23.995313 | 22.624670 | Interm. step, due to the fast rxns: [1] |
| 39 | 0.078 | 3.248265 | 23.641677 | 23.110059 | Interm. step, due to the fast rxns: [1] |
| 40 | 0.080 | 3.116512 | 23.298469 | 23.585019 | |
| 41 | 0.082 | 3.090374 | 22.859784 | 24.049842 | Interm. step, due to the fast rxns: [1] |
| 42 | 0.084 | 3.064236 | 22.433487 | 24.502277 | Interm. step, due to the fast rxns: [1] |
| 43 | 0.086 | 3.038098 | 22.019230 | 24.942672 | Interm. step, due to the fast rxns: [1] |
| 44 | 0.088 | 3.011960 | 21.616677 | 25.371363 | Interm. step, due to the fast rxns: [1] |
| 45 | 0.090 | 2.985822 | 21.225501 | 25.788677 | Interm. step, due to the fast rxns: [1] |
| 46 | 0.092 | 2.959684 | 20.845381 | 26.194935 | Interm. step, due to the fast rxns: [1] |
| 47 | 0.094 | 2.933546 | 20.476010 | 26.590444 | Interm. step, due to the fast rxns: [1] |
| 48 | 0.096 | 2.907408 | 20.117086 | 26.975507 | Interm. step, due to the fast rxns: [1] |
| 49 | 0.098 | 2.881270 | 19.768316 | 27.350415 | Interm. step, due to the fast rxns: [1] |
| 50 | 0.100 | 2.855132 | 19.429416 | 27.715453 | |
| 51 | 0.102 | 2.800545 | 19.128558 | 28.070897 | Interm. step, due to the fast rxns: [1] |
| 52 | 0.104 | 2.745959 | 18.836342 | 28.417698 | Interm. step, due to the fast rxns: [1] |
| 53 | 0.106 | 2.691373 | 18.552527 | 28.756100 | Interm. step, due to the fast rxns: [1] |
| 54 | 0.108 | 2.636787 | 18.276877 | 29.086336 | Interm. step, due to the fast rxns: [1] |
| 55 | 0.110 | 2.582201 | 18.009163 | 29.408636 | Interm. step, due to the fast rxns: [1] |
| 56 | 0.112 | 2.527614 | 17.749164 | 29.723221 | Interm. step, due to the fast rxns: [1] |
| 57 | 0.114 | 2.473028 | 17.496663 | 30.030308 | Interm. step, due to the fast rxns: [1] |
| 58 | 0.116 | 2.418442 | 17.251451 | 30.330107 | Interm. step, due to the fast rxns: [1] |
| 59 | 0.118 | 2.363856 | 17.013323 | 30.622821 | Interm. step, due to the fast rxns: [1] |
# Expand the last part
df.loc[60:]
| SYSTEM TIME | A | B | C | caption | |
|---|---|---|---|---|---|
| 60 | 0.120 | 2.309270 | 16.782080 | 30.908650 | |
| 61 | 0.122 | 2.282196 | 16.530018 | 31.187785 | Interm. step, due to the fast rxns: [1] |
| 62 | 0.124 | 2.255123 | 16.285122 | 31.459754 | Interm. step, due to the fast rxns: [1] |
| 63 | 0.126 | 2.228050 | 16.047192 | 31.724758 | Interm. step, due to the fast rxns: [1] |
| 64 | 0.128 | 2.200977 | 15.816031 | 31.982992 | Interm. step, due to the fast rxns: [1] |
| 65 | 0.130 | 2.173904 | 15.591452 | 32.234645 | Interm. step, due to the fast rxns: [1] |
| 66 | 0.132 | 2.146830 | 15.373269 | 32.479901 | Interm. step, due to the fast rxns: [1] |
| 67 | 0.134 | 2.119757 | 15.161303 | 32.718940 | Interm. step, due to the fast rxns: [1] |
| 68 | 0.136 | 2.092684 | 14.955381 | 32.951935 | Interm. step, due to the fast rxns: [1] |
| 69 | 0.138 | 2.065611 | 14.755333 | 33.179057 | Interm. step, due to the fast rxns: [1] |
| 70 | 0.140 | 2.038537 | 14.560994 | 33.400469 | |
| 71 | 0.142 | 2.010580 | 14.373089 | 33.616331 | Interm. step, due to the fast rxns: [1] |
| 72 | 0.144 | 1.982624 | 14.190557 | 33.826819 | Interm. step, due to the fast rxns: [1] |
| 73 | 0.146 | 1.954667 | 14.013248 | 34.032085 | Interm. step, due to the fast rxns: [1] |
| 74 | 0.148 | 1.926710 | 13.841015 | 34.232275 | Interm. step, due to the fast rxns: [1] |
| 75 | 0.150 | 1.898753 | 13.673717 | 34.427530 | Interm. step, due to the fast rxns: [1] |
| 76 | 0.152 | 1.870796 | 13.511215 | 34.617989 | Interm. step, due to the fast rxns: [1] |
| 77 | 0.154 | 1.842839 | 13.353374 | 34.803787 | Interm. step, due to the fast rxns: [1] |
| 78 | 0.156 | 1.814882 | 13.200065 | 34.985052 | Interm. step, due to the fast rxns: [1] |
| 79 | 0.158 | 1.786925 | 13.051161 | 35.161914 | Interm. step, due to the fast rxns: [1] |
| 80 | 0.160 | 1.758969 | 12.906537 | 35.334494 | |
| 81 | 0.180 | 1.572535 | 11.408782 | 37.018683 | |
| 82 | 0.200 | 1.385095 | 10.338861 | 38.276044 | |
| 83 | 0.220 | 1.266391 | 9.507280 | 39.226328 | |
| 84 | 0.240 | 1.166575 | 8.894402 | 39.939023 | |
| 85 | 0.260 | 1.096463 | 8.427419 | 40.476118 | |
| 86 | 0.280 | 1.041377 | 8.078969 | 40.879654 | |
| 87 | 0.300 | 1.001049 | 7.815530 | 41.183420 | |
| 88 | 0.320 | 0.970191 | 7.617998 | 41.411811 | |
| 89 | 0.340 | 0.947226 | 7.469116 | 41.583658 | |
| 90 | 0.360 | 0.929835 | 7.357265 | 41.712899 | |
| 91 | 0.380 | 0.916809 | 7.273064 | 41.810127 | |
| 92 | 0.400 | 0.906984 | 7.209759 | 41.883257 | last reaction step |
Notice:¶
the reaction proceeds in smaller steps in the earlier times (until t=0.160, in line 80), when the concentrations are changing much more rapidly
between lines 30 and 80 (time 0.060 to 0.160), only rection #1 (B <-> C) is regarded as fast-changing (based on the fast_threshold we specified in the simulation run); at earlier times, both reactions were regarded as fast-changing
"fast-changing" and "slow-changing" is NOT the same thing as "fast" and "slow" reaction kinetics. For example, reaction #1, though it has much slower kinetics than reaction #0, involves large concentration changes
after step 80, both reactions are regarded as slow-changing, and no more intermediate steps are used
Check the final equilibrium¶
# Verify that all the reactions have reached equilibrium
dynamics.is_in_equilibrium(tolerance=4)
A <-> B
Final concentrations: [B] = 7.21 ; [A] = 0.907
1. Ratio of reactant/product concentrations, adjusted for reaction orders: 7.94916
Formula used: [B] / [A]
2. Ratio of forward/reverse reaction rates: 8.0
Discrepancy between the two values: 0.6355 %
Reaction IS in equilibrium (within 4% tolerance)
B <-> C
Final concentrations: [C] = 41.88 ; [B] = 7.21
1. Ratio of reactant/product concentrations, adjusted for reaction orders: 5.80925
Formula used: [C] / [B]
2. Ratio of forward/reverse reaction rates: 6.0
Discrepancy between the two values: 3.179 %
Reaction IS in equilibrium (within 4% tolerance)
True
Plots of changes of concentration with time¶
dynamics.plot_curves(title="Coupled reactions A <-> B and B <-> C",
colors=['blue', 'red', 'green'])
dynamics.curve_intersection(t_start=0, t_end=0.05, var1="A", var2="B")
Min abs distance found at row: 6
(0.0111949581295586, 24.19287615172957)
dynamics.curve_intersection(t_start=0, t_end=0.05, var1="A", var2="C")
Min abs distance found at row: 16
(0.031791733478898146, 8.807902395796198)
dynamics.curve_intersection(t_start=0.05, t_end=0.1, var1="B", var2="C")
Min abs distance found at row: 40
(0.07929953388136385, 23.418671833563117)
# Verify that the stoichiometry is respected at every reaction step/substep (NOTE: it requires earlier activation of saving diagnostic data)
dynamics.stoichiometry_checker_entire_run()
True
Take a peek at the diagnostic data saved during the earlier reaction simulation¶
# This table contains a subset of a typical system history
dynamics.diagnostic_data_baselines.get()
| TIME | A | B | C | is_primary | primary_timestep | n_substeps | substep_number | caption | |
|---|---|---|---|---|---|---|---|---|---|
| 0 | 0.000 | 50.000000 | 0.000000 | 0.000000 | True | 0.02 | 10 | 0 | |
| 1 | 0.002 | 43.600000 | 6.400000 | 0.000000 | False | 0.02 | 10 | 1 | |
| 2 | 0.004 | 38.121600 | 11.724800 | 0.153600 | False | 0.02 | 10 | 2 | |
| 3 | 0.006 | 33.429632 | 16.135987 | 0.434381 | False | 0.02 | 10 | 3 | |
| 4 | 0.008 | 29.408815 | 19.771278 | 0.819907 | False | 0.02 | 10 | 4 | |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 88 | 0.320 | 0.970191 | 7.617998 | 41.411811 | True | 0.02 | 10 | 0 | |
| 89 | 0.340 | 0.947226 | 7.469116 | 41.583658 | True | 0.02 | 10 | 0 | |
| 90 | 0.360 | 0.929835 | 7.357265 | 41.712899 | True | 0.02 | 10 | 0 | |
| 91 | 0.380 | 0.916809 | 7.273064 | 41.810127 | True | 0.02 | 10 | 0 | |
| 92 | 0.400 | 0.906984 | 7.209759 | 41.883257 | True | 0.02 | 10 | 0 |
93 rows × 9 columns
# Concentration increments due to reaction 0 (A <-> B)
# Note that [C] is not affected
dynamics.get_diagnostic_data(rxn_index=0)
Reaction: A <-> B
| TIME | Delta A | Delta B | Delta C | reaction | substep | time_subdivision | delta_time | caption | |
|---|---|---|---|---|---|---|---|---|---|
| 0 | 0.000 | -6.400000 | 6.400000 | 0.0 | 0 | 0 | 10 | 0.002 | |
| 1 | 0.002 | -5.478400 | 5.478400 | 0.0 | 0 | 1 | 10 | 0.002 | |
| 2 | 0.004 | -4.691968 | 4.691968 | 0.0 | 0 | 2 | 10 | 0.002 | |
| 3 | 0.006 | -4.020817 | 4.020817 | 0.0 | 0 | 3 | 10 | 0.002 | |
| 4 | 0.008 | -3.447988 | 3.447988 | 0.0 | 0 | 4 | 10 | 0.002 | |
| 5 | 0.010 | -2.959017 | 2.959017 | 0.0 | 0 | 5 | 10 | 0.002 | |
| 6 | 0.012 | -2.541571 | 2.541571 | 0.0 | 0 | 6 | 10 | 0.002 | |
| 7 | 0.014 | -2.185132 | 2.185132 | 0.0 | 0 | 7 | 10 | 0.002 | |
| 8 | 0.016 | -1.880728 | 1.880728 | 0.0 | 0 | 8 | 10 | 0.002 | |
| 9 | 0.018 | -1.620710 | 1.620710 | 0.0 | 0 | 9 | 10 | 0.002 | |
| 10 | 0.020 | -1.398555 | 1.398555 | 0.0 | 0 | 0 | 10 | 0.002 | |
| 11 | 0.022 | -1.208698 | 1.208698 | 0.0 | 0 | 1 | 10 | 0.002 | |
| 12 | 0.024 | -1.046394 | 1.046394 | 0.0 | 0 | 2 | 10 | 0.002 | |
| 13 | 0.026 | -0.907597 | 0.907597 | 0.0 | 0 | 3 | 10 | 0.002 | |
| 14 | 0.028 | -0.788857 | 0.788857 | 0.0 | 0 | 4 | 10 | 0.002 | |
| 15 | 0.030 | -0.687228 | 0.687228 | 0.0 | 0 | 5 | 10 | 0.002 | |
| 16 | 0.032 | -0.600202 | 0.600202 | 0.0 | 0 | 6 | 10 | 0.002 | |
| 17 | 0.034 | -0.525637 | 0.525637 | 0.0 | 0 | 7 | 10 | 0.002 | |
| 18 | 0.036 | -0.461708 | 0.461708 | 0.0 | 0 | 8 | 10 | 0.002 | |
| 19 | 0.038 | -0.406858 | 0.406858 | 0.0 | 0 | 9 | 10 | 0.002 | |
| 20 | 0.040 | -0.359757 | 0.359757 | 0.0 | 0 | 0 | 10 | 0.002 | |
| 21 | 0.042 | -0.319273 | 0.319273 | 0.0 | 0 | 1 | 10 | 0.002 | |
| 22 | 0.044 | -0.284441 | 0.284441 | 0.0 | 0 | 2 | 10 | 0.002 | |
| 23 | 0.046 | -0.254434 | 0.254434 | 0.0 | 0 | 3 | 10 | 0.002 | |
| 24 | 0.048 | -0.228551 | 0.228551 | 0.0 | 0 | 4 | 10 | 0.002 | |
| 25 | 0.050 | -0.206192 | 0.206192 | 0.0 | 0 | 5 | 10 | 0.002 | |
| 26 | 0.052 | -0.186845 | 0.186845 | 0.0 | 0 | 6 | 10 | 0.002 | |
| 27 | 0.054 | -0.170073 | 0.170073 | 0.0 | 0 | 7 | 10 | 0.002 | |
| 28 | 0.056 | -0.155505 | 0.155505 | 0.0 | 0 | 8 | 10 | 0.002 | |
| 29 | 0.058 | -0.142822 | 0.142822 | 0.0 | 0 | 9 | 10 | 0.002 | |
| 30 | 0.060 | -1.317528 | 1.317528 | 0.0 | 0 | 0 | 1 | 0.020 | |
| 31 | 0.080 | -0.261380 | 0.261380 | 0.0 | 0 | 0 | 1 | 0.020 | |
| 32 | 0.100 | -0.545862 | 0.545862 | 0.0 | 0 | 0 | 1 | 0.020 | |
| 33 | 0.120 | -0.270732 | 0.270732 | 0.0 | 0 | 0 | 1 | 0.020 | |
| 34 | 0.140 | -0.279569 | 0.279569 | 0.0 | 0 | 0 | 1 | 0.020 | |
| 35 | 0.160 | -0.186434 | 0.186434 | 0.0 | 0 | 0 | 1 | 0.020 | |
| 36 | 0.180 | -0.187439 | 0.187439 | 0.0 | 0 | 0 | 1 | 0.020 | |
| 37 | 0.200 | -0.118704 | 0.118704 | 0.0 | 0 | 0 | 1 | 0.020 | |
| 38 | 0.220 | -0.099816 | 0.099816 | 0.0 | 0 | 0 | 1 | 0.020 | |
| 39 | 0.240 | -0.070112 | 0.070112 | 0.0 | 0 | 0 | 1 | 0.020 | |
| 40 | 0.260 | -0.055086 | 0.055086 | 0.0 | 0 | 0 | 1 | 0.020 | |
| 41 | 0.280 | -0.040328 | 0.040328 | 0.0 | 0 | 0 | 1 | 0.020 | |
| 42 | 0.300 | -0.030858 | 0.030858 | 0.0 | 0 | 0 | 1 | 0.020 | |
| 43 | 0.320 | -0.022965 | 0.022965 | 0.0 | 0 | 0 | 1 | 0.020 | |
| 44 | 0.340 | -0.017391 | 0.017391 | 0.0 | 0 | 0 | 1 | 0.020 | |
| 45 | 0.360 | -0.013027 | 0.013027 | 0.0 | 0 | 0 | 1 | 0.020 | |
| 46 | 0.380 | -0.009825 | 0.009825 | 0.0 | 0 | 0 | 1 | 0.020 |
# Concentration increments due to reaction 1 (B <-> C)
# Note that [A] is not affected
dynamics.get_diagnostic_data(rxn_index=1)
Reaction: B <-> C
| TIME | Delta A | Delta B | Delta C | reaction | substep | time_subdivision | delta_time | caption | |
|---|---|---|---|---|---|---|---|---|---|
| 0 | 0.000 | 0.0 | 0.000000 | 0.000000 | 1 | 0 | 10 | 0.002 | |
| 1 | 0.002 | 0.0 | -0.153600 | 0.153600 | 1 | 1 | 10 | 0.002 | |
| 2 | 0.004 | 0.0 | -0.280781 | 0.280781 | 1 | 2 | 10 | 0.002 | |
| 3 | 0.006 | 0.0 | -0.385526 | 0.385526 | 1 | 3 | 10 | 0.002 | |
| 4 | 0.008 | 0.0 | -0.471231 | 0.471231 | 1 | 4 | 10 | 0.002 | |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 87 | 0.300 | 0.0 | -0.228390 | 0.228390 | 1 | 0 | 1 | 0.020 | |
| 88 | 0.320 | 0.0 | -0.171847 | 0.171847 | 1 | 0 | 1 | 0.020 | |
| 89 | 0.340 | 0.0 | -0.129241 | 0.129241 | 1 | 0 | 1 | 0.020 | |
| 90 | 0.360 | 0.0 | -0.097228 | 0.097228 | 1 | 0 | 1 | 0.020 | |
| 91 | 0.380 | 0.0 | -0.073130 | 0.073130 | 1 | 0 | 1 | 0.020 |
92 rows × 9 columns
# Expand the last part of the above table
dynamics.get_diagnostic_data(rxn_index=1).loc[60:]
Reaction: B <-> C
| TIME | Delta A | Delta B | Delta C | reaction | substep | time_subdivision | delta_time | caption | |
|---|---|---|---|---|---|---|---|---|---|
| 60 | 0.120 | 0.0 | -0.279135 | 0.279135 | 1 | 0 | 10 | 0.002 | |
| 61 | 0.122 | 0.0 | -0.271969 | 0.271969 | 1 | 1 | 10 | 0.002 | |
| 62 | 0.124 | 0.0 | -0.265004 | 0.265004 | 1 | 2 | 10 | 0.002 | |
| 63 | 0.126 | 0.0 | -0.258234 | 0.258234 | 1 | 3 | 10 | 0.002 | |
| 64 | 0.128 | 0.0 | -0.251653 | 0.251653 | 1 | 4 | 10 | 0.002 | |
| 65 | 0.130 | 0.0 | -0.245256 | 0.245256 | 1 | 5 | 10 | 0.002 | |
| 66 | 0.132 | 0.0 | -0.239039 | 0.239039 | 1 | 6 | 10 | 0.002 | |
| 67 | 0.134 | 0.0 | -0.232996 | 0.232996 | 1 | 7 | 10 | 0.002 | |
| 68 | 0.136 | 0.0 | -0.227121 | 0.227121 | 1 | 8 | 10 | 0.002 | |
| 69 | 0.138 | 0.0 | -0.221412 | 0.221412 | 1 | 9 | 10 | 0.002 | |
| 70 | 0.140 | 0.0 | -0.215862 | 0.215862 | 1 | 0 | 10 | 0.002 | |
| 71 | 0.142 | 0.0 | -0.210489 | 0.210489 | 1 | 1 | 10 | 0.002 | |
| 72 | 0.144 | 0.0 | -0.205266 | 0.205266 | 1 | 2 | 10 | 0.002 | |
| 73 | 0.146 | 0.0 | -0.200190 | 0.200190 | 1 | 3 | 10 | 0.002 | |
| 74 | 0.148 | 0.0 | -0.195255 | 0.195255 | 1 | 4 | 10 | 0.002 | |
| 75 | 0.150 | 0.0 | -0.190459 | 0.190459 | 1 | 5 | 10 | 0.002 | |
| 76 | 0.152 | 0.0 | -0.185797 | 0.185797 | 1 | 6 | 10 | 0.002 | |
| 77 | 0.154 | 0.0 | -0.181266 | 0.181266 | 1 | 7 | 10 | 0.002 | |
| 78 | 0.156 | 0.0 | -0.176861 | 0.176861 | 1 | 8 | 10 | 0.002 | |
| 79 | 0.158 | 0.0 | -0.172580 | 0.172580 | 1 | 9 | 10 | 0.002 | |
| 80 | 0.160 | 0.0 | -1.684189 | 1.684189 | 1 | 0 | 1 | 0.020 | |
| 81 | 0.180 | 0.0 | -1.257360 | 1.257360 | 1 | 0 | 1 | 0.020 | |
| 82 | 0.200 | 0.0 | -0.950285 | 0.950285 | 1 | 0 | 1 | 0.020 | |
| 83 | 0.220 | 0.0 | -0.712694 | 0.712694 | 1 | 0 | 1 | 0.020 | |
| 84 | 0.240 | 0.0 | -0.537096 | 0.537096 | 1 | 0 | 1 | 0.020 | |
| 85 | 0.260 | 0.0 | -0.403536 | 0.403536 | 1 | 0 | 1 | 0.020 | |
| 86 | 0.280 | 0.0 | -0.303766 | 0.303766 | 1 | 0 | 1 | 0.020 | |
| 87 | 0.300 | 0.0 | -0.228390 | 0.228390 | 1 | 0 | 1 | 0.020 | |
| 88 | 0.320 | 0.0 | -0.171847 | 0.171847 | 1 | 0 | 1 | 0.020 | |
| 89 | 0.340 | 0.0 | -0.129241 | 0.129241 | 1 | 0 | 1 | 0.020 | |
| 90 | 0.360 | 0.0 | -0.097228 | 0.097228 | 1 | 0 | 1 | 0.020 | |
| 91 | 0.380 | 0.0 | -0.073130 | 0.073130 | 1 | 0 | 1 | 0.020 |
Provide a detailed explanation of all the steps/substeps of the reactions, from the saved diagnostic data¶
# dynamics.explain_reactions() # Uncomment if desired
Re-run with very small constanst steps¶
We'll use constant steps of size 0.0005, which is 1/4 of the smallest steps (the "substep" size) previously used in the variable-step run
dynamics2 = ReactionDynamics(reaction_data=chem_data)
dynamics2.set_conc([50., 0, 0.], snapshot=True)
dynamics2.single_compartment_react(time_step=0.0005, reaction_duration=0.4,
snapshots={"initial_caption": "1st reaction step",
"final_caption": "last reaction step"},
)
800 total step(s) taken
fig = px.line(data_frame=dynamics2.get_history(), x="SYSTEM TIME", y=["A", "B", "C"],
title="Coupled reactions A <-> B and B <-> C (run with constant steps)",
color_discrete_sequence = ['blue', 'red', 'green'],
labels={"value":"concentration", "variable":"Chemical"})
fig.show()
dynamics2.curve_intersection(t_start=0, t_end=0.05, var1="A", var2="B")
Min abs distance found at row: 24
(0.011934802021672202, 24.03028826511689)
dynamics2.curve_intersection(t_start=0, t_end=0.05, var1="A", var2="C")
Min abs distance found at row: 65
(0.03259520587890533, 9.081194286771336)
dynamics2.curve_intersection(t_start=0.05, t_end=0.1, var1="B", var2="C")
Min abs distance found at row: 159
(0.07932921476313338, 23.238745930657448)
df2 = dynamics2.get_history()
df2
| SYSTEM TIME | A | B | C | caption | |
|---|---|---|---|---|---|
| 0 | 0.0000 | 50.000000 | 0.000000 | 0.000000 | Initial state |
| 1 | 0.0005 | 48.400000 | 1.600000 | 0.000000 | 1st reaction step |
| 2 | 0.0010 | 46.857600 | 3.132800 | 0.009600 | |
| 3 | 0.0015 | 45.370688 | 4.600925 | 0.028387 | |
| 4 | 0.0020 | 43.937230 | 6.006806 | 0.055964 | |
| ... | ... | ... | ... | ... | ... |
| 796 | 0.3980 | 0.925494 | 7.329151 | 41.745354 | |
| 797 | 0.3985 | 0.925195 | 7.327221 | 41.747584 | |
| 798 | 0.3990 | 0.924898 | 7.325303 | 41.749800 | |
| 799 | 0.3995 | 0.924602 | 7.323396 | 41.752002 | |
| 800 | 0.4000 | 0.924309 | 7.321501 | 41.754190 | last reaction step |
801 rows × 5 columns
Notice that we now did 801 steps - vs. the 93 of the earlier variable-resolution run!¶
Let's compare some entries with the coarser previous variable-time run¶
At time t=0.002:
df.loc[1]
SYSTEM TIME 0.002 A 43.6 B 6.4 C 0.0 caption Interm. step, due to the fast rxns: [0, 1] Name: 1, dtype: object
df2.loc[4] # High-precision result from fine fixed-resolution run
SYSTEM TIME 0.002 A 43.93723 B 6.006806 C 0.055964 caption Name: 4, dtype: object
old = df.loc[1][['SYSTEM TIME', 'A', 'B', 'C']].to_numpy().astype(float)
old
array([2.00e-03, 4.36e+01, 6.40e+00, 0.00e+00])
new = df2.loc[4][['SYSTEM TIME', 'A', 'B', 'C']].to_numpy().astype(float)
new
array([2.00000000e-03, 4.39372297e+01, 6.00680596e+00, 5.59643616e-02])
new - old
array([ 0. , 0.33722968, -0.39319404, 0.05596436])
At time t=0.032, when [A] and [C] are almost equal:
df.loc[16]
SYSTEM TIME 0.032 A 8.736339 B 32.378083 C 8.885578 caption Interm. step, due to the fast rxns: [0, 1] Name: 16, dtype: object
df2.loc[64] # High-precision result from fine fixed-resolution run
SYSTEM TIME 0.032 A 9.282079 B 31.85364 C 8.864282 caption Name: 64, dtype: object
At time t=0.14:
df.loc[70]
SYSTEM TIME 0.14 A 2.038537 B 14.560994 C 33.400469 caption Name: 70, dtype: object
df2.loc[280] # High-precision result from fine fixed-resolution run
SYSTEM TIME 0.14 A 2.070976 B 14.710086 C 33.218938 caption Name: 280, dtype: object
At time t=0.26:
df.loc[85]
SYSTEM TIME 0.26 A 1.096463 B 8.427419 C 40.476118 caption Name: 85, dtype: object
df2.loc[520] # High-precision result from fine fixed-resolution run
SYSTEM TIME 0.26 A 1.145609 B 8.749175 C 40.105216 caption Name: 520, dtype: object
Let's compare the plots of [B] from the earlier (variable-step) run, and the latest (high-precision fixed-step) one¶
# Earlier run
fig1 = px.line(data_frame=dynamics.get_history(), x="SYSTEM TIME", y=["B"],
color_discrete_sequence = ['red'],
labels={"value":"concentration", "variable":"Variable-step run"})
fig1.show()
# Latest run (High-precision result from fine fixed-resolution run)
fig2 = px.line(data_frame=dynamics2.get_history(), x="SYSTEM TIME", y=["B"],
color_discrete_sequence = ['orange'],
labels={"value":"concentration", "variable":"Fine fixed-step run"})
fig2.show()
import plotly.graph_objects as go
all_fig = go.Figure(data=fig1.data + fig2.data) # Note that the + is concatenating lists
all_fig.update_layout(title="The 2 runs contrasted")
all_fig['data'][0]['name']="B (variable steps)"
all_fig['data'][1]['name']="B (fixed high precision)"
all_fig.show()